TWOMBLI uses some of these metrics and adds others more pertinent to the study of matrix fibres. Ridge detection is an algorithm for detecting the filamentous structure of an image in an unbiased manner and Anamorf is a plugin for quantifying such structures. TWOMBLI relies heavily on existing ImageJ plugins Ridge Detection and AnaMorf ( 16, 17), providing an end-to-end pipeline for users. TWOMBLI exists as a user-friendly ImageJ plugin that can be downloaded with detailed documentation from. The aim of TWOMBLI is to quantify matrix patterns in an ECM image by deriving a range of metrics, which can then be analysed in conjunction with clinical data if desired. To this end, we have created the ImageJ macro plugin TWOMBLI, which stands for The Workflow Of Matrix BioLogy Informatics. There is a need for an end-to-end pipeline for quantifying ECM patterns, which is automated and easy-to-use on versatile data sets. This makes collating a broad range of metrics for analysis both challenging and time-consuming. Furthermore, each tool typically only generates metrics relating to one aspect of ECM organisation.
#Imagej quantify he staining manual#
A number of studies have attempted to quantify additional matrix metrics, but these typically require heavy manual intervention ( 14, 15). OrientationJ is an automated ImageJ plugin that is able to create vector fields and perform directional analysis on fibres, but this does not lend itself to quantifying overall matrix patterns. A pipeline already exists in MATLAB for quantifying alignment of ECM ( 12), and a further extension of this work enables alignment to be related to the tumour margin ( 13). One commonly used metric is matrix fibre alignment, which is known to be a promoter of cancer cell invasion ( 9, 10, 11). These range from simple abundance and area measurements to more complex textural features such as grey level co-occurrence matrices ( 7), which measures texture by analysing the variation in pixel intensity value, and box-counting fractal dimension ( 8), which measures how an image is sampled in “boxes” of decreasing size. Numerous metrics have already been applied to such ECM images. Furthermore, the transition of isotropic and curved matrices into more linear aligned matrices is associated with aggressive cancers. Highly aligned and linear matrix fibres are well suited to resisting tensile stresses in the direction of alignment, and can also serve as migration routes for cells. The generation of metrics that describe ECM pattern could lead to insights into a wide range of fields, ranging from experimentalists interested in cell migration and remodelling of matrices to clinicians researching conditions such as cancer and fibrosis.ĭiverse ECM patterns are observed in both normal and pathological tissue. The architecture of the ECM has lately been the subject of renewed focus ( 4, 5, 6) however, standardised quantification of ECM patterns, particularly in pathological decision-making is lacking. Changes in the ECM are central to the ageing process, the ECM is re-built and remodelled in response to tissue damage, and it is further altered in pathologies such as cancer.
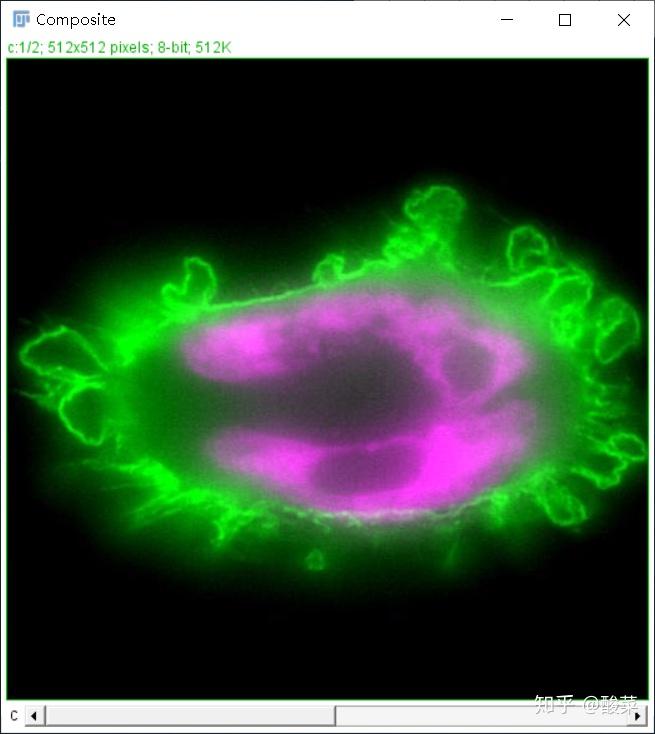
The ECM provides support and structure to multicellular organisms and also guides the migration of cells ( 1, 2), including leukocytes engaged in immune surveillance ( 3). Here we present an overview of the pipeline together with examples from a wide range of contexts where matrix patterns are generated. Although developed with the extracellular matrix in mind, TWOMBLI is versatile and can be applied to vascular and cytoskeletal networks. TWOMBLI can be downloaded from together with detailed documentation and tutorial video. TWOMBLI is designed to be quick, versatile and easy-to-use particularly for non-computational scientists. This pipeline includes metrics of matrix alignment, length, branching, end points, gaps, fractal dimension, curvature, and the distribution of fibre thickness. To this end, we have developed an ImageJ plugin called TWOMBLI, which stands for The Workflow Of Matrix BioLogy Informatics. Thus, an automated pipeline that simultaneously quantifies a broad range of metrics and enables a comprehensive description of varied matrix patterns is needed. However, most current tools for quantitative analysis focus on a single aspect of matrix patterning. Diverse extracellular matrix patterns are observed in both normal and pathological tissue.
